Improving Chemical Understanding of LLMs via SMILES Parsing
EMNLP 2025
KAIST, Yonsei University
Overview
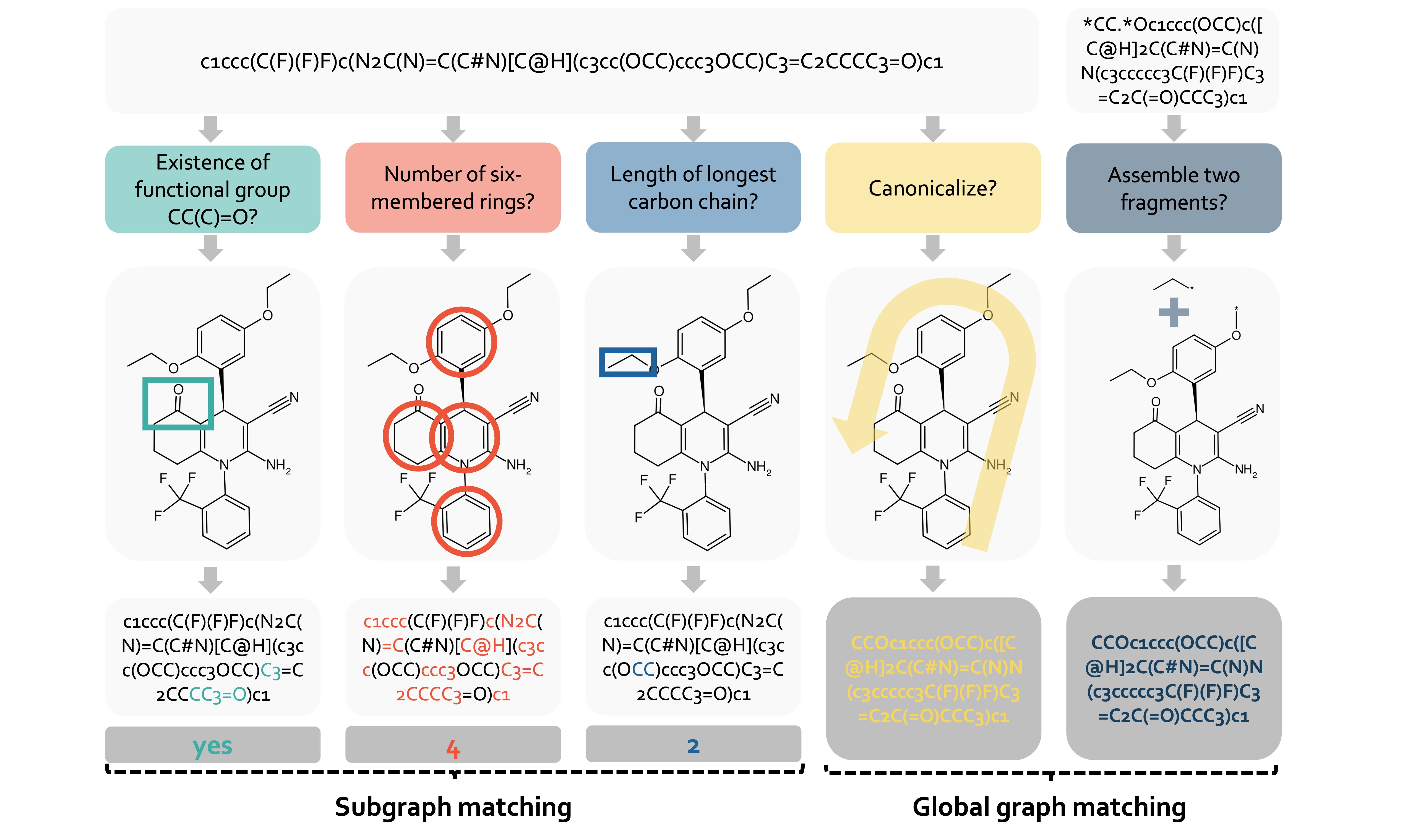
We address limitations in how large language models interpret molecular structures encoded in SMILES format. We introduce CLEANMOL, a framework converting SMILES parsing into structured tasks designed to enhance graph-level molecular comprehension. The approach spans from subgraph to global graph matching with adaptive difficulty scoring. Results demonstrate improved structural understanding and competitive performance on the Mol-Instructions benchmark.
BibTeX
@inproceedings{jang-etal-2025-improving,
title = "Improving Chemical Understanding of {LLM}s via {SMILES} Parsing",
author = "Jang, Yunhui and
Kim, Jaehyung and
Ahn, Sungsoo",
editor = "Christodoulopoulos, Christos and
Chakraborty, Tanmoy and
Rose, Carolyn and
Peng, Violet",
booktitle = "Proceedings of the 2025 Conference on Empirical Methods in Natural Language Processing",
month = nov,
year = "2025",
address = "Suzhou, China",
publisher = "Association for Computational Linguistics",
url = "https://aclanthology.org/2025.emnlp-main.791/",
doi = "10.18653/v1/2025.emnlp-main.791",
pages = "15683--15698",
ISBN = "979-8-89176-332-6"
}